Ferrington Lab Group
Research Projects
Determining the mechanism linking mitochondrial defects in the retinal pigment epithelium with the CFH risk allele for age-related macular degeneration
 While the pathogenesis of age-related macular degeneration (AMD) is not completely elucidated, genome-wide association studies of AMD patients have identified polymorphisms in 34 high-risk loci involving different biological pathways, including complement, cell survival, lipid transport and processing, extracellular matrix remodeling and angiogenesis. One of the most prevalent AMD-associated single-nucleotide polymorphisms (SNP) occurs in the gene encoding complement factor H (CFH). CFH is a negative regulator of the alternative complement pathway and protects against inappropriate activation that can cause chronic inflammation. The SNP in CFH (rs1061170), where a tyrosine is substituted by a histidine at position 402 (Y402H) in the protein, is found in ~50% of AMD patients. How this single amino acid change contributes to AMD pathogenesis is unclear. Our published work, showing significant mtDNA damage and lower mitochondrial function in the retinal pigment epithelium (RPE) of AMD donors with the CFH high-risk SNP, suggest mitochondrial (mt) defects are associated with the presence of the CFH high risk variant. The hypothesis is that the presence of the CFH high risk allele alters mt function and the response to metabolic and oxidative stressors.
While the pathogenesis of age-related macular degeneration (AMD) is not completely elucidated, genome-wide association studies of AMD patients have identified polymorphisms in 34 high-risk loci involving different biological pathways, including complement, cell survival, lipid transport and processing, extracellular matrix remodeling and angiogenesis. One of the most prevalent AMD-associated single-nucleotide polymorphisms (SNP) occurs in the gene encoding complement factor H (CFH). CFH is a negative regulator of the alternative complement pathway and protects against inappropriate activation that can cause chronic inflammation. The SNP in CFH (rs1061170), where a tyrosine is substituted by a histidine at position 402 (Y402H) in the protein, is found in ~50% of AMD patients. How this single amino acid change contributes to AMD pathogenesis is unclear. Our published work, showing significant mtDNA damage and lower mitochondrial function in the retinal pigment epithelium (RPE) of AMD donors with the CFH high-risk SNP, suggest mitochondrial (mt) defects are associated with the presence of the CFH high risk variant. The hypothesis is that the presence of the CFH high risk allele alters mt function and the response to metabolic and oxidative stressors.
Our approach integrates classical reductionist investigations, coupled with discovery-driven analyses (proteomics and metabolomics), to reveal novel information aimed at explaining the ~50% prevalence of the CFH high-risk allele in patients with AMD. Model systems include primary RPE and iPSC-derived RPE cultured from individuals that have been phenotyped for disease severity and genotyped for the CFH risk allele. Reciprocal isogenic iPSC-RPE lines generated from parent iPSCs using CRISPR-Cas9 gene editing provide the most direct test of the genotype-specific effects on metabolic and functional outcomes since the cells are genetically identical except for a single amino acid change in CFH.
This ongoing project, funded by an R01 grant from NEI/NIH, aims to elucidate the mechanism responsible for development of AMD in the population of patients harboring the CFH risk allele. This information is critical for development of new therapies that target the primary defect.
Funding: National Institutes of Health/National Eye Institute 2 R01 EY028554-05
Collaborators: Production of iPSC-RPE and isogenic lines – Drs. James Dutton and Beau Webber, Donor grading – Dr. Sandra Montezuma, University of Minnesota. Proteomic analysis – Dr. Jun Qu, University of Buffalo. Metabolomic analysis – Dr. Jianhai Du, University of West Virginia.
Determine the sites of failure in processes associated with maintenance of mitochondrial homeostasis
Our previous investigations support the hypothesis that mitochondrial damage and dysfunction is an early event in AMD pathology. The overall hypothesis is that damaged mitochondria accumulate in RPE from AMD donors due to defects in one or more of the processes associated with mitochondrial homeostasis. This project involves measuring the major processes of mitochondrial homeostasis, including biogenesis, fission/fusion, and autophagic removal of damaged mitochondria. The studies are performed in human donor RPE using a variety of assays (i.e., immunoblotting, immunofluorescence) and advanced imaging techniques using fluorescent proteins targeted to the mitochondria (mKeima, GFP-mito) and organelle-specific fluorescent dyes (mito-tracker, lysotracker).
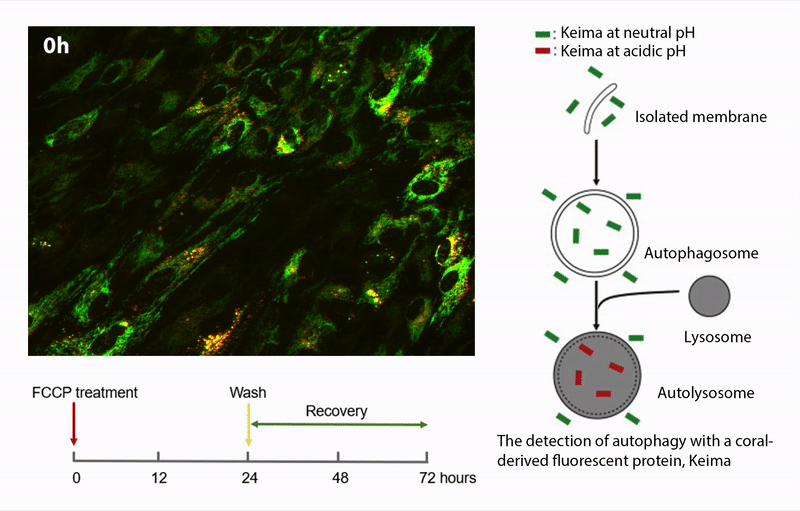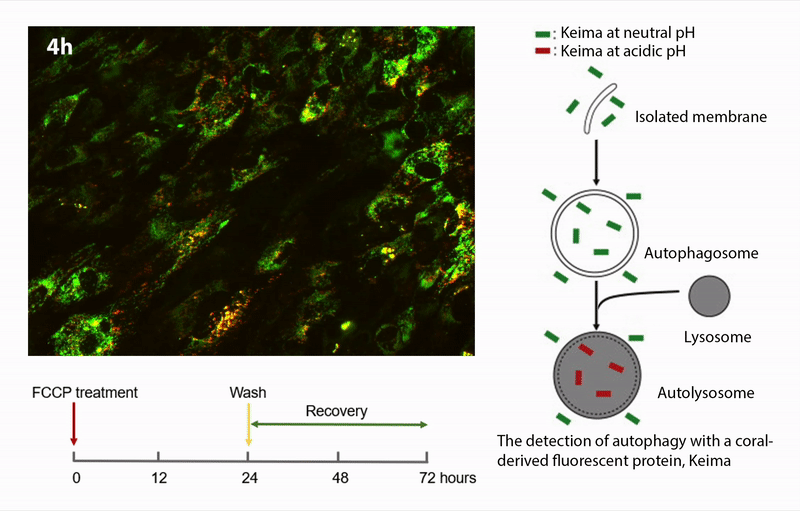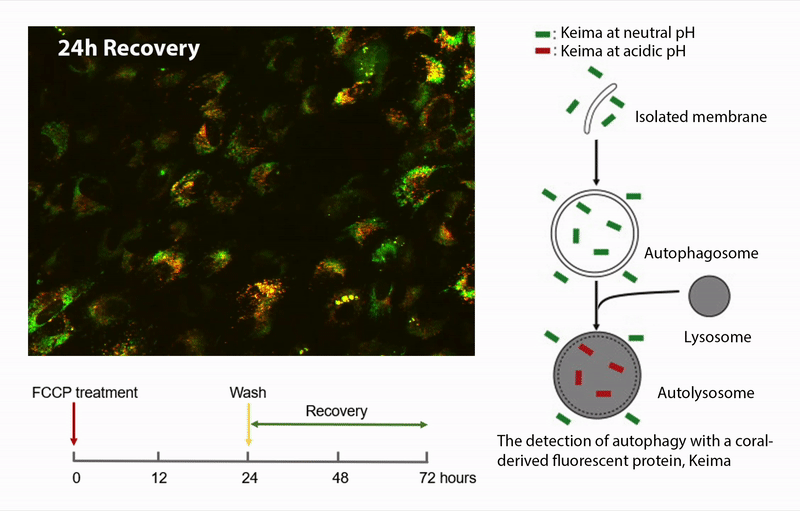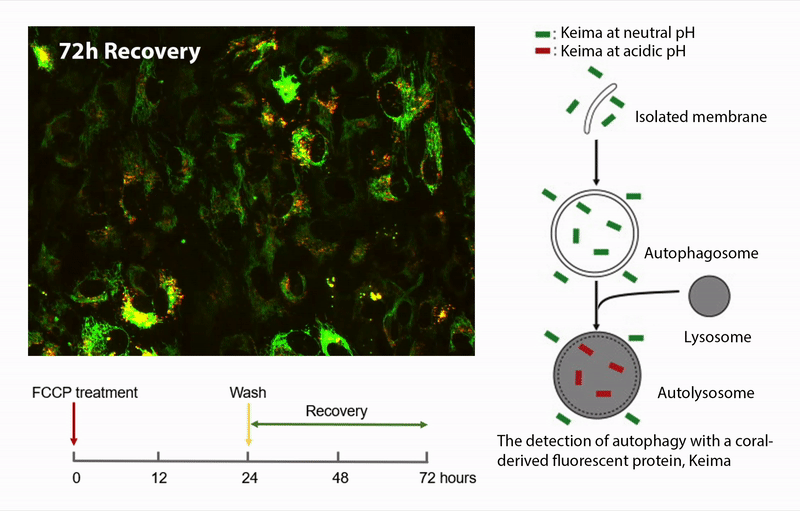
Funding: National Institutes of Health/National Eye Institute 2 R01 EY028554-05
Genetic and epigenetic analyses identify novel regulators and contributors of AMD
In recent years, large-scale genetic studies have helped to identify several key molecules as well as non-coding genomic regions in pathogenesis of AMD. However, we do not understand the complex networks through which candidates operate or the functional consequences of genetic variation on quantitative traits. Using retinas from human donors graded for the presence and severity of AMD, the Swaroop laboratory is performing an integrated analysis of the transcriptome (RNA sequencing), genome (whole-genome sequencing/genotyping), and epigenome (DNA Methylation and histone modifications). The goal is to generate quantitative trait loci maps of human retina to elucidate gene-environment interactions, identify causal variants and delineate their contribution to disease pathology.
Funding: National Eye Institute Intramural Research Program, Lindsay Family Foundation, and the Doheny Eye Institute.
Collaborators: Donor grading – Dr. Sandra Montezuma, University of Minnesota. Genetic and Epigenetic Analyses – Dr. Anand Swaroop, National Eye Institute, Bethesda, MD.
Determining the mechanism behind the transition from healthy aging to early age-related macular degeneration
This project asks, “what are the molecular events in the RPE that precipitate the transition from healthy aging to early AMD?” To answer this question, we use RPE tissue from human donor eyes graded for the presence and severity of AMD using the Minnesota Grading System (MGS). Donor tissue is used to perform a highly sensitive and comprehensive proteomic analysis to define which proteins are altered in RPE tissue isolated from healthy aged (MGS1, no AMD) donors and compared with RPE from donors with early AMD (MGS2) using a novel IonStar quantitative strategy. Our initial analysis identified multiple biological pathways that have changed during the transition to early-stage disease, indicating potential mechanisms that initiate the pathological switch. This new information, which identifies the pathological events that tip the balance from healthy aging to early disease, is critical for the development of targeted therapies designed to eliminate the progression to blindness experienced by a large percentage of our elderly population.
Funding: Winston and Maxine Wallin Neuroscience Discovery Fund, Lindsay Family Foundation and the Doheny Eye Institute.
Collaborators: Proteomic analysis – Dr. Jun Qu, University of Buffalo. Donor grading – Dr. Sandra Montezuma, University of Minnesota

Identify drugs that protect or improve RPE mitochondrial function
There are currently no effective treatments for dry AMD, which includes ~80% of all AMD patients. Our prior investigations show mitochondrial damage occurs at an early stage of AMD. We proposed treatments that protect the mitochondria or help boost function could stop or attenuate AMD progression. This project utilizes RPE differentiated from induced pluripotent stem cells (iPSCs) to screen panels of FDA-approved drugs with known beneficial effects on the mitochondria. The goal is to identify drugs with efficacious effects on RPE mitochondria to provide preclinical “proof of concept” for future in vivo research.
Funding: Lindsay Family Foundation and the Doheny Eye Institute.
Collaborators: Production of iPSC-RPE – Drs. James Dutton, Grading donors- Sandra Montezuma, University of Minnesota.